
⚠️ Unexpected Maintenance Notice
The service will undergo unexpected maintenance on Thursday, March 26th, 2026 at 11:00 AM (CET).
The system may become temporarily unavailable during this time. Running jobs may be interrupted and could be lost.
We strongly recommend avoiding job submissions shortly before the maintenance window.
We apologize for the inconvenience and thank you for your understanding.
BUSCA (Bologna Unified Subcellular Component Annotator) is a web-server for predicting protein subcellular localization. BUSCA integrates different tools to predict localization-related protein features (DeepSig, TPpred3, PredGPI, BetAware and ENSEMBLE3.0) as well as tools for discriminating subcellular localization of both globular and membrane proteins (BaCelLo, MemLoci and SChloro).
For details see: Savojardo, C, Martelli, PL, Fariselli, P, Profiti, G, Casadio, R (2018) BUSCA: an integrative web server to predict subcellular localization of proteins. Nucleid Acids Research, 46(W1), W459-W466.
BUSCA is able to predict:
- 16 compartments for plant proteins
- 9 compartments for animal or fungi proteins
- 4 compartments for Gram-negative bacterial proteins
- 3 compartments for Gram-positive bacterial proteins
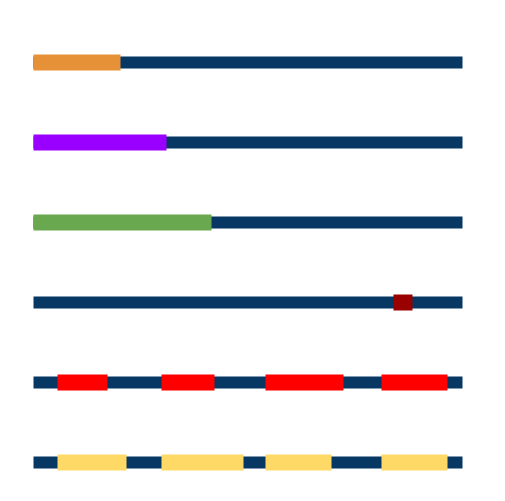
BUSCA provides residue-level annotation of the following features:
- Secretory signal peptides
- Organelle-targeting peptides (mitochondrial and chloroplastic)
- GPI-anchors
- Alpha and beta transmembrane regions